Mohan and Pearl have just had published a paper ‘Graphical Models for Processing Missing Data’ (open access pre-print here, journal version here). It’s a great read, and no doubt contains lots of useful developments (I’m still working my way through the paper). But something strikes me as somewhat troubling about their missing at random definition. Years ago when working with colleagues on using directed acyclic graphs to encode missing data assumptions, we struggled to see how MAR monotone dropout, as might occur in a longitudinal study, could be encoded in a DAG. In this post I will try and see whether MAR monotone dropout is classified as MAR according the definitions of Mohan and Pearl.
Suppose we have a study which intends to measure individuals on some outcome at baseline (Y0) and two follow-up time points (Y1,Y2). Assume everyone has Y0, the outcome at baseline, measured. Suppose that some individuals then dropout from the study (and hence have Y1 and Y2 missing), with their dropout decision made dependent on Y0. Among those who don’t dropout before Y1 is measured, some dropout before Y2 is measured. Suppose that among those who have Y1 measured, the decision of whether to dropout is made dependent solely based on Y1. Under these assumptions, the missing data mechanism satisfies the MAR assumption (see Robins and Gill 1997), in the sense that the probability of a particular pattern of variables being observed only depends on the values of variables which are observed in that pattern.
As far as I understand, the main distinction with Mohan and Pearl’s missingness graphs are the inclusion of proxy variables. There is one for each partially observed variable, and it is a function of the corresponding variable’s missingness indicator and the underlying variable in question. The proxy variable equals the value of the variable in question when that variable is observed, and otherwise takes some dummy value (Mohan and Pearl label this value m). I believe the m-graph for the above MAR monotone dropout scenario is as follows:

For simplicity I have assumed that Y0 doesn’t have a direct effect on Y2. Here R1 and R2 are the binary missingness indicators of Y1 and Y2 respectively. Y_1 and Y_2 are the proxy variables for Y1 and Y2 (Mohan and Pearl use the variable names post-fixed with a *, but my DAG drawing package can’t handle this character!). Note that whether or not Y2 is observed depends only on Y_1, the proxy version of Y1. This encodes the fact that we have assumed that missingness in Y2 only depends on Y1 in those who didn’t dropout before Y1 was to be measured. For those who dropped out before Y1 could be measured, Y_1 indicates this by taking value m, and R2 is then deterministically 1 (meaning Y2 is missing according to Mohan and Pearl’s notation).
Now we can use this m-graph to check MAR according to Mohan and Pearl’s definition. They define V_O and V_M as the sets of variables that are fully observed and partially observed respectively. In our example, V_O=Y0, which we assume is fully observed, while V_M={Y1,Y2}, since each of these variables is partially observed. According to my understanding of Mohan and Pearl’s definition in their Section 2.2, to check MAR (or v-MAR, as Mohan and Pearl label it) we have to check if each missingness indicator variable (R1 and R2 here) is independent of V_M conditional on V_O=Y0.
Conditional on Y0, R1 is independent of {Y1,Y2} because the path via Y0 is blocked (since we are conditioning on it) and the paths via Y_1 and Y_2 are also blocked because these are colliders. So far, so good. Now we must check if R2 is independent of {Y1,Y2} conditional on Y0. According to the graph, we cannot read off that this is true, since there is an open path from R2 to Y1 via the proxy variable Y_1.
I may well have made a mistake in the above reasoning, in which case it would be great if someone could add a comment explaining it. Otherwise this seems a little problematic. We have a quite standard well established situation where the implication of the classical definition of MAR is widely understood. In the context of longitudinal studies, if we are willing to assume MAR we can use likelihood based methods such as mixed models to handle the missing data. If we follow Mohan and Pearl’s method and definition in this case, we apparently conclude we have MNAR, which at least if one looked at the historical literature, would imply a mixed model analysis would be biased (which we know it isn’t, under these assumptions).
In an earlier 2013 paper, Mohan, Pearl and Tian, where as far as I know Mohan and Pearl first proposed m-graphs, they wrote:
This graph-based interpretation uses slightly stronger assumptions than Rubin’s, with the advantage that the user can comprehend, encode and communicate the assumptions that determine the classification of the problem.
https://ftp.cs.ucla.edu/pub/stat_ser/r410.pdf
This sentence would seem to support my suspicion that there is indeed a quite important difference between classical definitions of MAR (see Seaman et al 2013 for a careful review) and Mohan and Pearl’s definition, which if users are not aware of, could cause confusion, particularly if they are familiar with the older definition(s) of MAR and its implications for validity of certain types of statistical analysis. But there is also a significant probability that I have misunderstood Mohan and Pearl or made a mistake in the above!
Postscript
Karthika Mohan kindly replied to me on Twitter and confirmed my understanding above is correct:
1/4 You are absolutely right. We would classify this problem as MNAR and, as we stress in our paper, MAR (Rubin,1976) differs from v-MAR (section 2.2). Moreover, while most practitioners would tend to give up on hearing the word "MNAR," we rejoice the challenge of handling such
2/4 problems with the general methods developed. The m-graph in your blog is interesting, but it contains arrows emanating from proxy variables. We'd represent the same missingness process equivalently without such arrows, as shown below. From the standpoint of physics, proxy
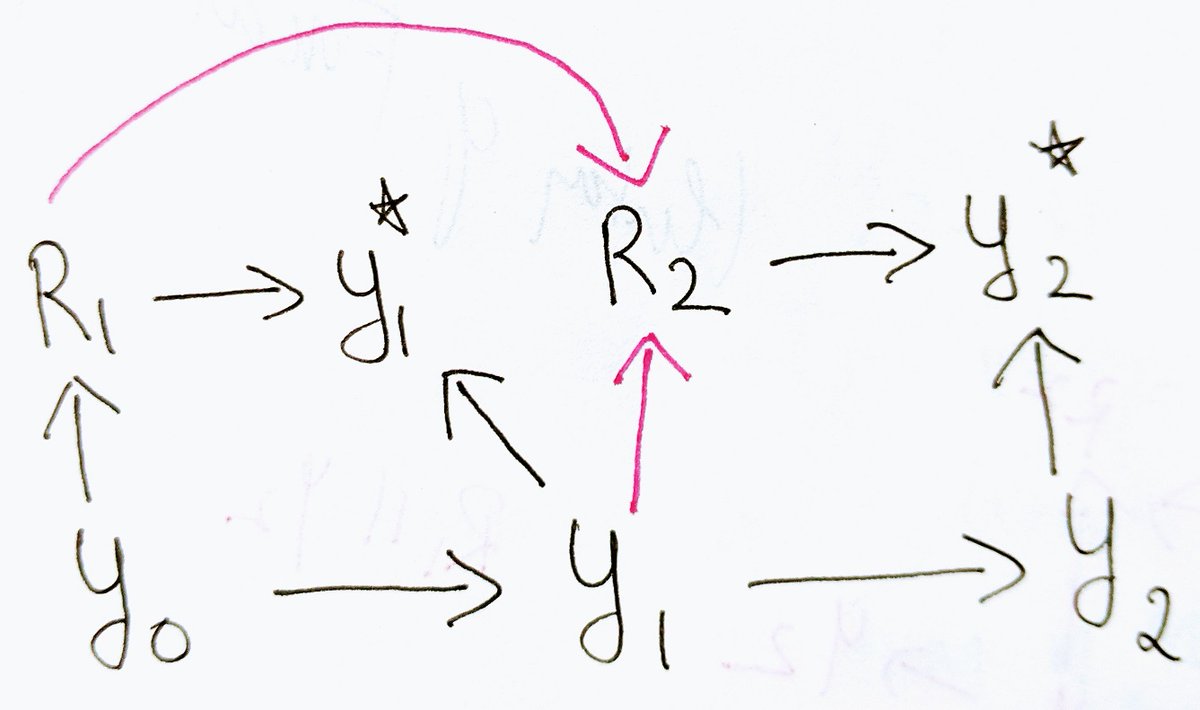
3/4 variables are passive manifestations of physical events and do not cause other events. In this m-graph, joint distribution is recoverable using Sequential Factorization (section 3.1). Consequently, every statistical quantity and every identifiable causal effect is recoverable
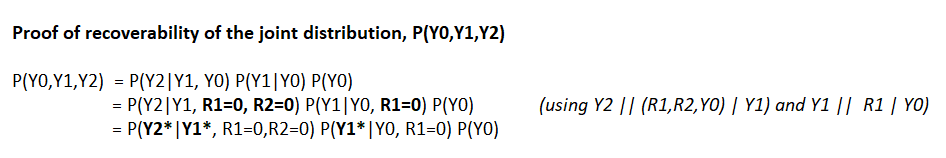
4/4 Bonus for graph-friendly researchers – Does the model have testable implications? You bet it does: e.g. Y0||R2| Y1,R1=0. See rules for detecting testable implications (section 4.1). Can we get these results from the graph-less literature? I doubt it and, honestly, why labor?
Originally tweeted by Karthika Mohan (@Carthica) on January 18, 2021.
MAR monotone dropout does not satisfy their v-MAR definition, and would therefore be classified as MNAR by their framework. Thus while ‘classical MAR’ is not the same as Mohan and Pearl’s v-MAR, by implication ‘classical MNAR’ is not the same as Mohan and Pearl’s MNAR (they don’t write v-MNAR for some reason in the paper).
This is not my area of expertise but I think you need to think of how information is accumulating over time. (If you know about filtration, I think that is what you have). Y2 is not MAR given only Y0. Since if Y1 is none missing whether or not Y2 is missing depends on the value of Y1. So too in Mohan & Pearl V_o and V_m change with time.
It may be helpful to think about (right) censoring in survival analysis. When is the actual failure time MAR. If you set as a counting process rather than a simple failure time then you see how MAR is conditional on the history.
Thanks Peter. Yes, Y2 is not MAR given only Y0. The fact that missingness in Y2 only depends on Y1 when Y1 is observed is encoded in the DAG by the fact that R2 depends on Y_1, the proxy for Y1 that takes the value of Y1 iff Y1 is observed.
You say V_O and V_M can change with time, but I see nothing in the setup or definitions of Section 2 of their paper that state this. V_O and V_M are simply defined as the set of variables in the original DAG which are fully observed and partially observed respectively.
I think you’re both right. Causal DAGs should be dynamic in exactly the sense that Peter describes, sharply distinguishing between past and future. Filtrations seem to me a very natural way to achieve this.
But Jonathan is right that MAR (and also, apparently, v-MAR) are static conditions, and it is not always easy to see whether a plausible, dynamic, causal setup implies the static MAR. We’ve had two recent attempts:
1. https://academic.oup.com/biomet/article/104/2/317/3804413
(Uses causal DAGs to demonstrate ignorable observation in continuous time, without appealing to notions of missing data.)
2. https://academic.oup.com/biomet/advance-article/doi/10.1093/biomet/asab002/6128501
(Identifies missing at random with adaptedness to a particular filtration, and extensible to continuous time.)